Genomic sequence might still be the most underexploited resource in understanding biology. Even though genomic sequence is widely used in research as a reference to make measurements about the genome or detect variants, the information encoded in the sequence, the biology encoded in the genome, is still rarely extracted as a major driver for scientific discoveries.
Understanding genomic sequence is a problem that is challenging with traditional biology approaches, but it is almost a perfect problem to address for machine learning and AI. We now have complete genome sequences and hundreds of thousands of genome-wide assays measuring sequence-dependent activities across the genome, from transcription factor binding to 3D genome organization to transcription. Capturing the link between sequence and these activities represents an unusually rich opportunity for computation-driven discovery.
Our group currently focuses on three major directions. The first direction is prediction, which involves predicting sequence activities directly from DNA. The second is understanding, where we develop models and interpretations that expose the underlying sequence-based mechanisms of regulatory activity. The third is design, where we reverse the prediction task and generate sequences with desired activities at different scales. These three directions reinforce one another and together shape the lab's current research program.
Recent research projects
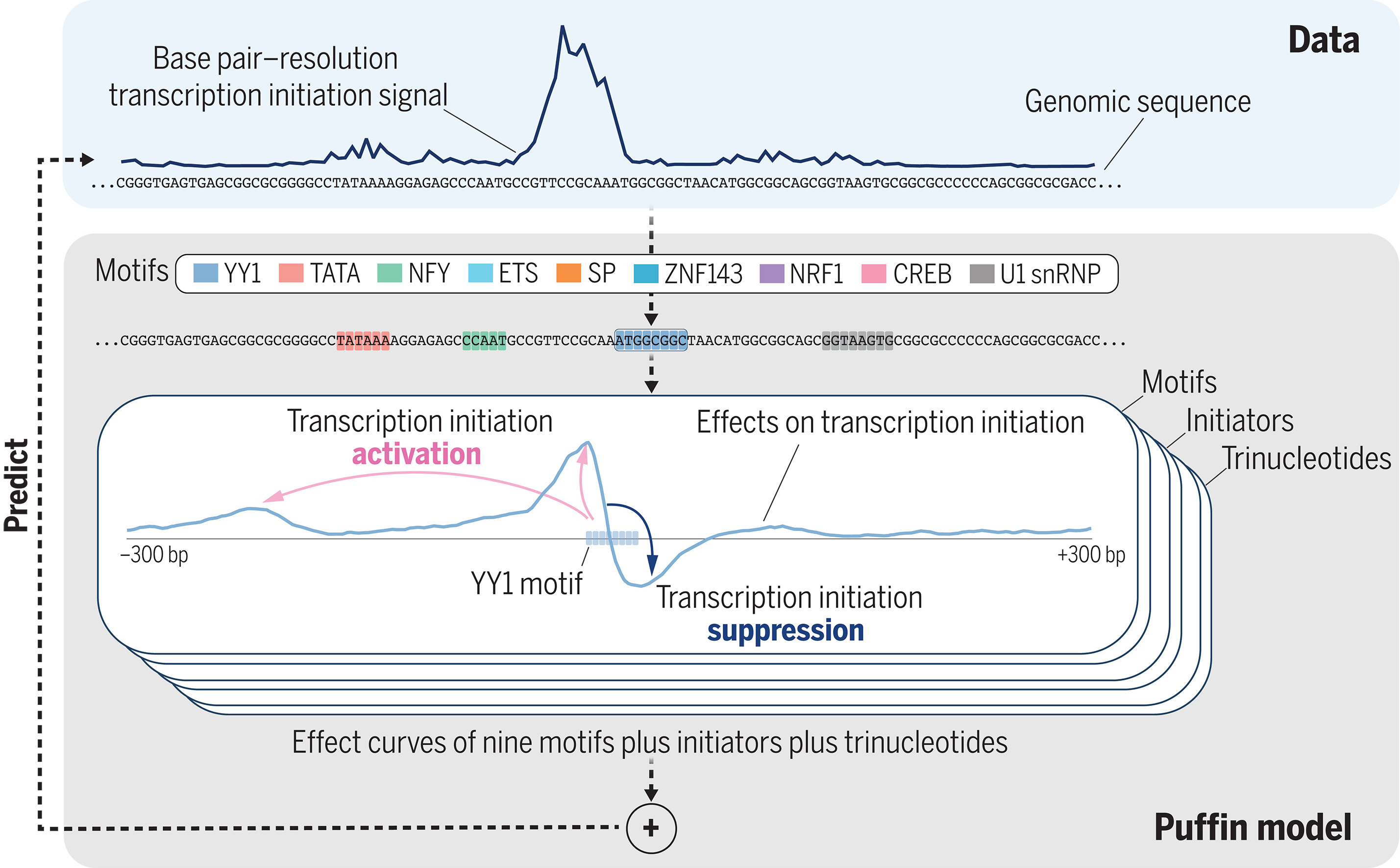
Despite the central role of promoters in transcription initiation, promoter logic in the human genome remained poorly understood. Puffin is an explainable machine learning model that dissects how transcription initiation depends on sequence, identifies a compact set of motifs and rules that explain most promoters, and offers a unified view of promoter sequence and function.
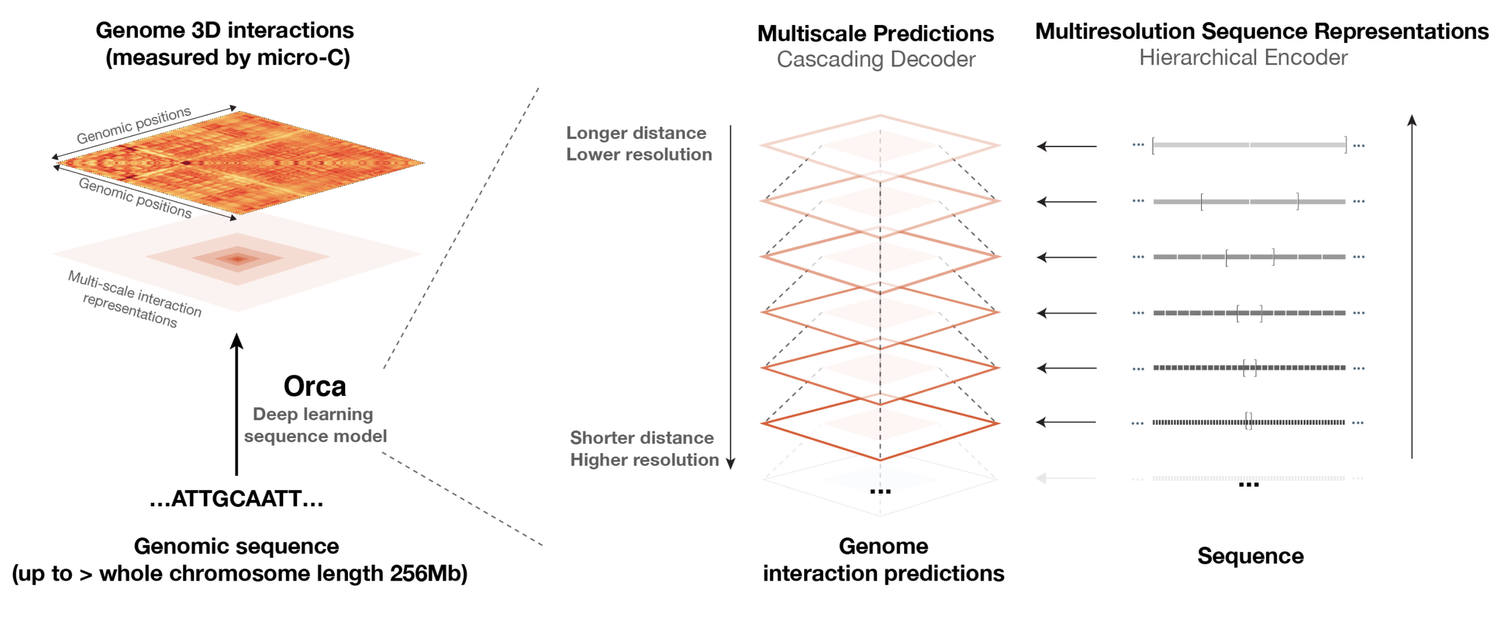
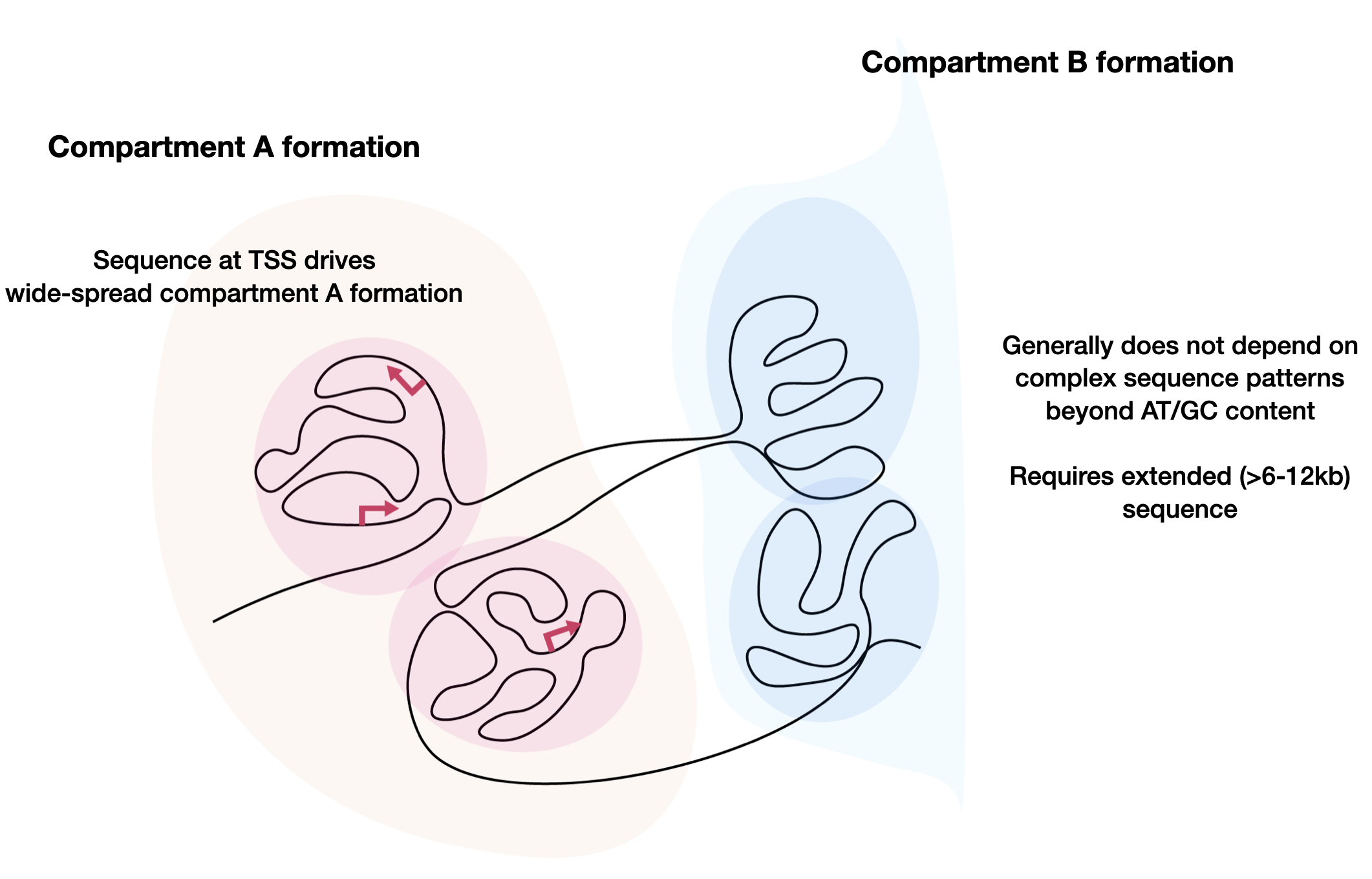
Orca is a sequence-based deep learning model for predicting 3D genome architecture from kilobase to whole-chromosome scale, including the impact of structural variants. The model also points to a putative sequence basis for chromatin compartment formation.
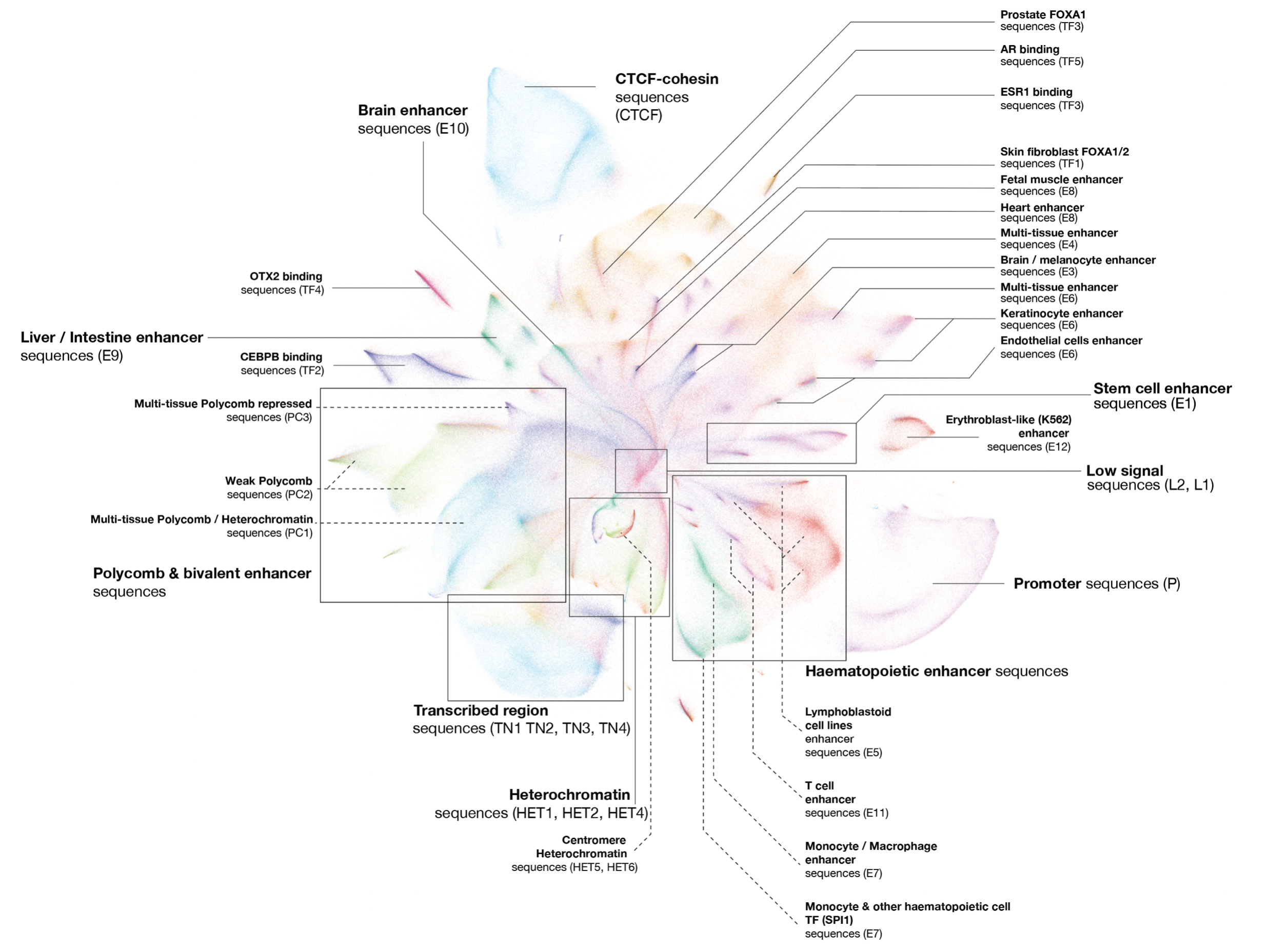
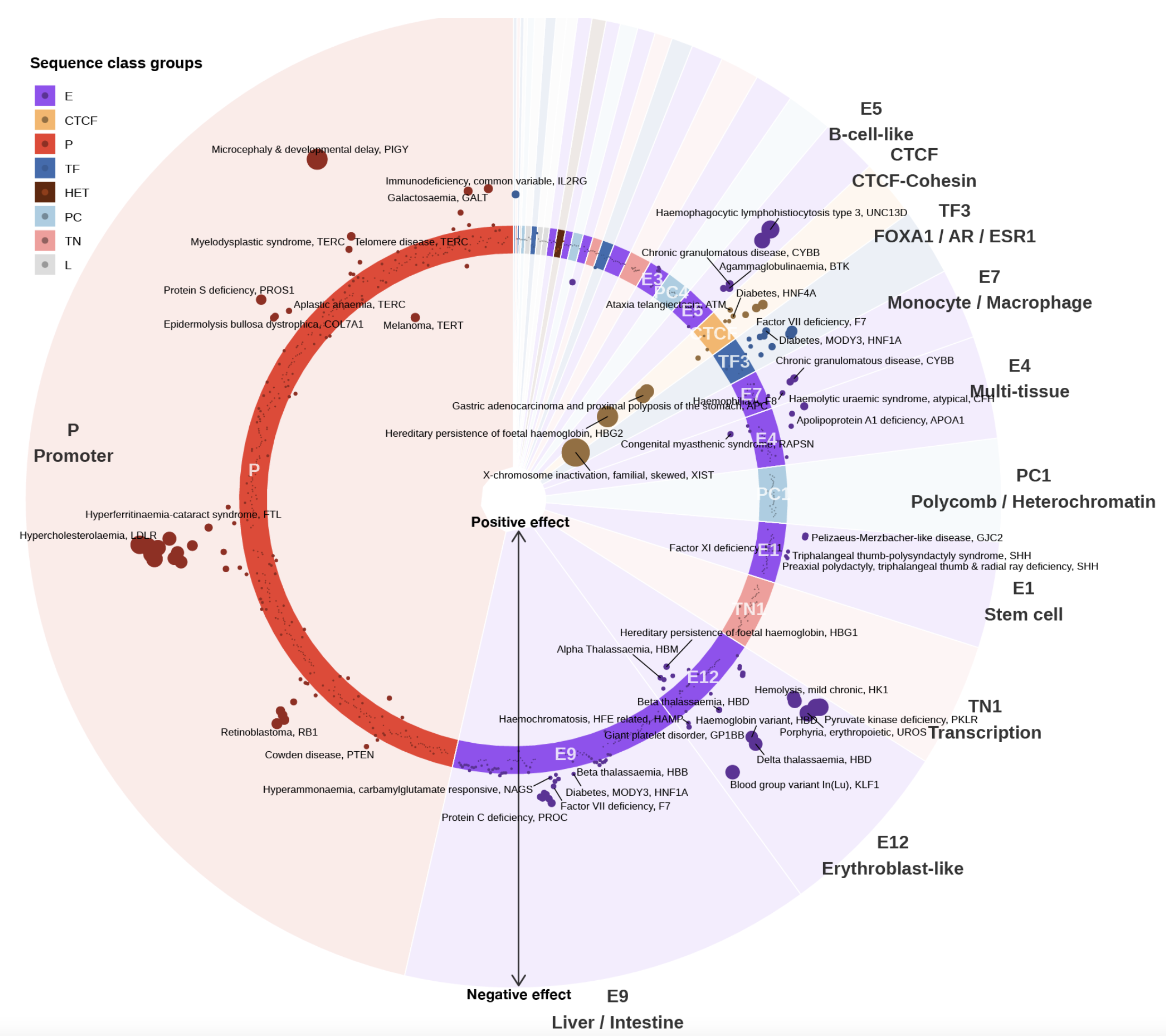
Sei integrates human genetics data with sequence information to discover the regulatory basis of traits and diseases. It learns a vocabulary of regulatory sequence classes using a deep learning model trained across more than 21,000 chromatin profiles from over 1,300 cell lines and tissues.